Phage Patience's Novel Capsid Structure Reveals New Insights into Viral Evolution and Genome Accommodation
April 3, 2025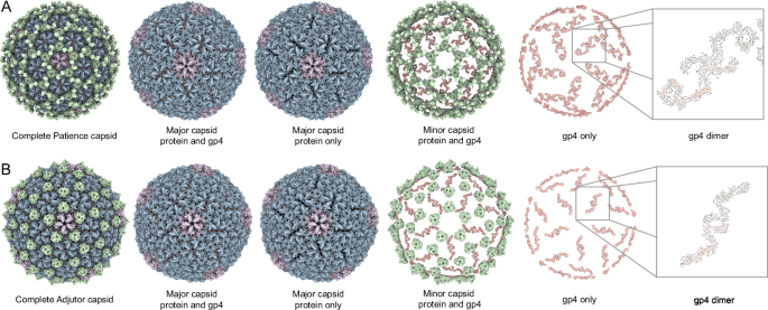
Recent research has focused on actinobacteriophages, analyzing a collection of 4,000 annotated genomes through cryo-electron microscopy (cryo-EM) to explore capsid structures and their evolutionary relationships.
Accessory proteins, such as gp4 found in Patience and its homolog in Adjutor, play a crucial role in stabilizing the split hexamers, facilitating increased genome sizes without enlarging the capsid.
Among these, the phage Patience stands out with its T = 7 icosahedral capsid, which can accommodate a 70.5 kb genome, exceeding typical size limitations due to an increase in internal volume.
Additionally, mathematical principles have been identified that predict which icosahedral capsids can utilize similar stabilization mechanisms, revealing that capsids with T-numbers not centered on global 3-fold axes can accommodate split hexamers.
Tailed bacteriophage capsids can accommodate genome lengths ranging from 10 to 750 kilobase pairs, supported by a sophisticated maturation process that stabilizes the capsid during assembly.
The study delves into the stabilization mechanisms of tailed bacteriophage capsids, which protect their genomes while adapting to various sizes of viral DNA.
This maturation process involves scaffolding proteins and major capsid proteins (MCPs) forming a procapsid that undergoes conformational changes during DNA packaging, resulting in a stable, mature capsid.
The HK97-fold structure of MCPs is essential, featuring conserved domains that enable significant conformational changes, allowing the capsid to adapt to larger genomes.
Patience's capsid is characterized by split hexamers, a novel structural feature that enhances genomic accommodation, previously unobserved in mature capsids.
The research indicates that incremental changes in the length of accessory proteins can help adapt capsids to larger genomic lengths, providing valuable insights into viral evolution.
Summary based on 1 source
