Revolutionary Spatial Transcriptomics Method Unveiled: Breakthrough in Gene Expression Mapping by Broad Institute Team
April 3, 2025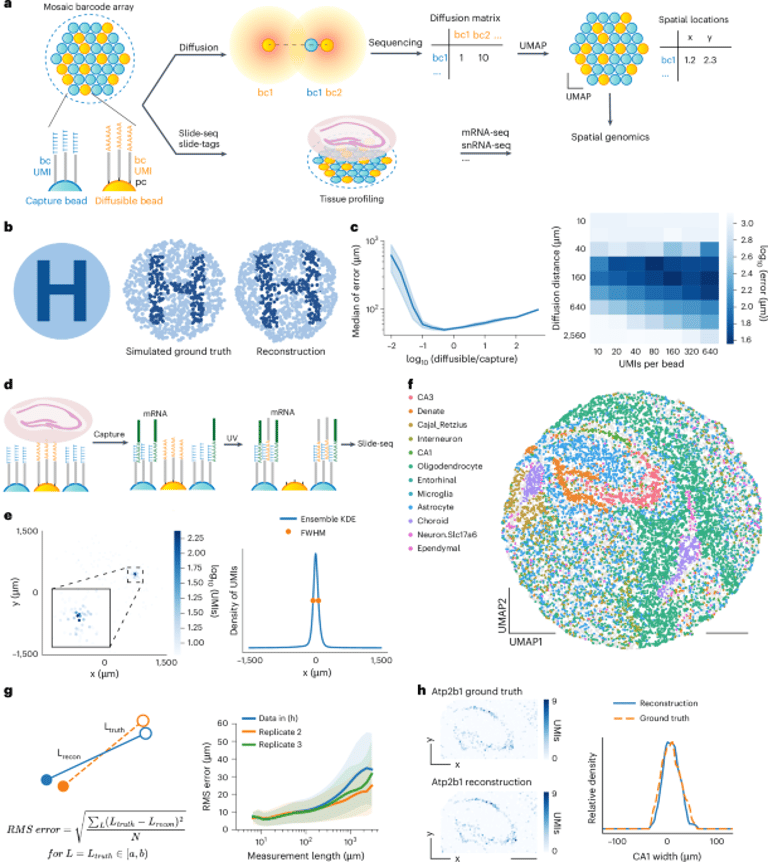
A new method for scalable spatial transcriptomics has been developed, enabling high-resolution gene expression mapping within tissues.
Researchers from the Broad Institute of Harvard and MIT, led by Chenlei Hu, created a protocol that enhances the accuracy of spatial transcriptomics.
This research highlights the significance of scalable techniques in spatial transcriptomics, which are crucial for analyzing complex biological systems and understanding disease mechanisms.
Among the significant findings, the study demonstrated accurate reconstruction of gene expression patterns, offering valuable insights into cellular organization, particularly in the mouse hippocampus.
The method utilizes a computational framework that reconstructs the spatial organization of gene expression from data gathered through Slide-seq and Slide-tags technologies.
Key contributors to this study included Mehdi Borji, Giovanni J. Marrero, and Evan Z. Macosko, who played vital roles in experiments and computational analysis.
The research received support from several institutions, including the National Institutes of Health and the New York Stem Cell Foundation.
The article also provides supplementary data, including extended figures that illustrate simulation results and performance metrics of the reconstruction method.
The authors acknowledged the contributions of various individuals who assisted with tissue handling, library preparation, and discussions regarding computational algorithms.
Summary based on 1 source